88 figure panels, 1 straightforward message:
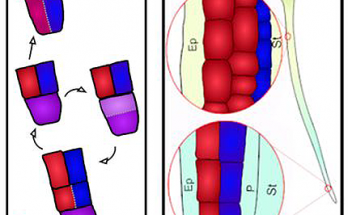
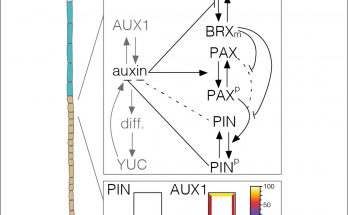
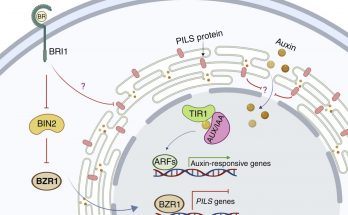
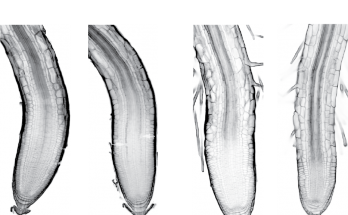
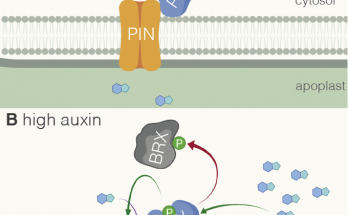
Phosphosites in the linker region of BRX family proteins determine whether or not their association with the plasma membrane association is sensitive to auxin. Congratulations to Sam and co-workers for …
88 figure panels, 1 straightforward message: Read More