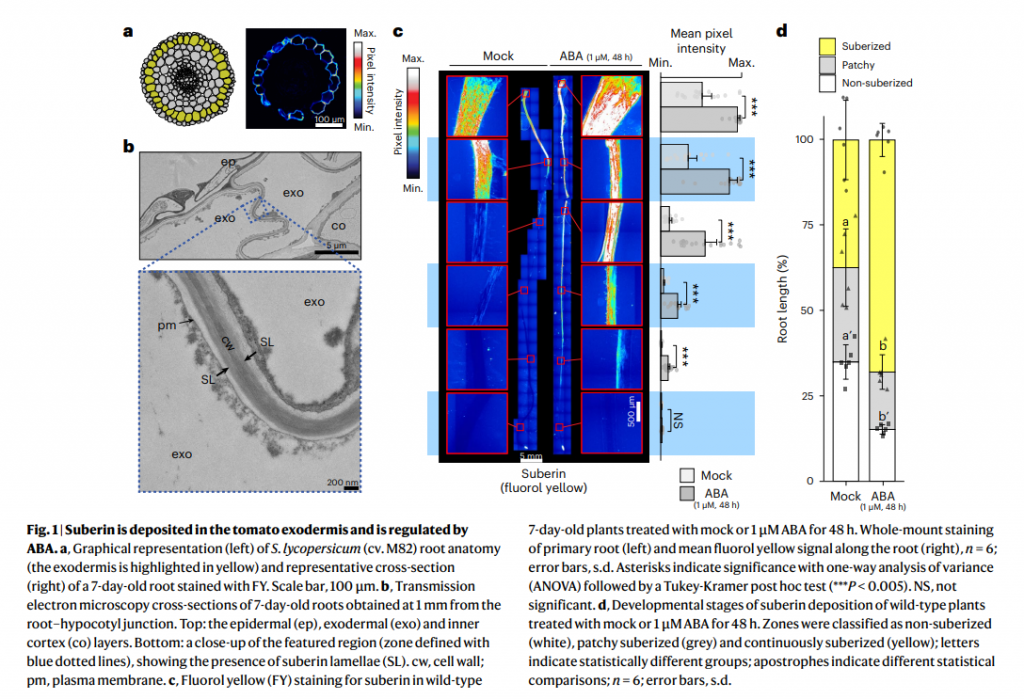
Here a new article in Nature Plant done with the help of Damien De Bellis from the EMF

As a core facility dedicated to service at the Faculty of Biology and Medicine, we serve in priority investigators from all departments of the UNIL. We are providing full service, training on all our instruments and autonomous access to trained users. External users are welcome to contact us to discuss opportunities.
Here a new article in Nature Plant done with the help of Damien De Bellis from the EMF

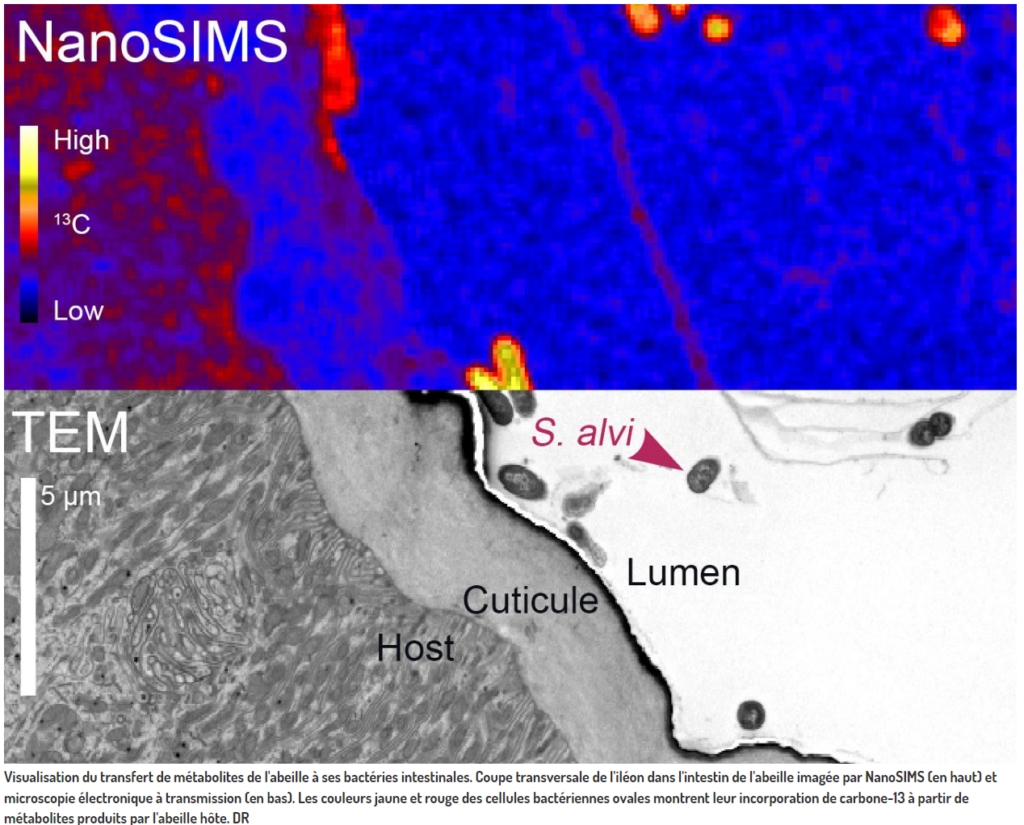
We are proud to have contributed to the this study published in « Nature Microbiology ». Jean Daraspe has received the bee guts and prepare them for EM and NanoSIMs. Then he took TEM images of fields of view where the bacteriae are visible in gut before giving the samples to the group of A Meibom that has performed NanoSIMS on them… Many thanks to the group of Pr Engel to recognize the contribution of the EMF to this study and Congratulations for this pioneered work!
It made it to CQFD on the RTS radio for the francophones!
Between Christmas and New Year, we received 10 huge boxes, one being 1200 kg … The new Talos L120 is in his definitive new room and will be installed next week!



